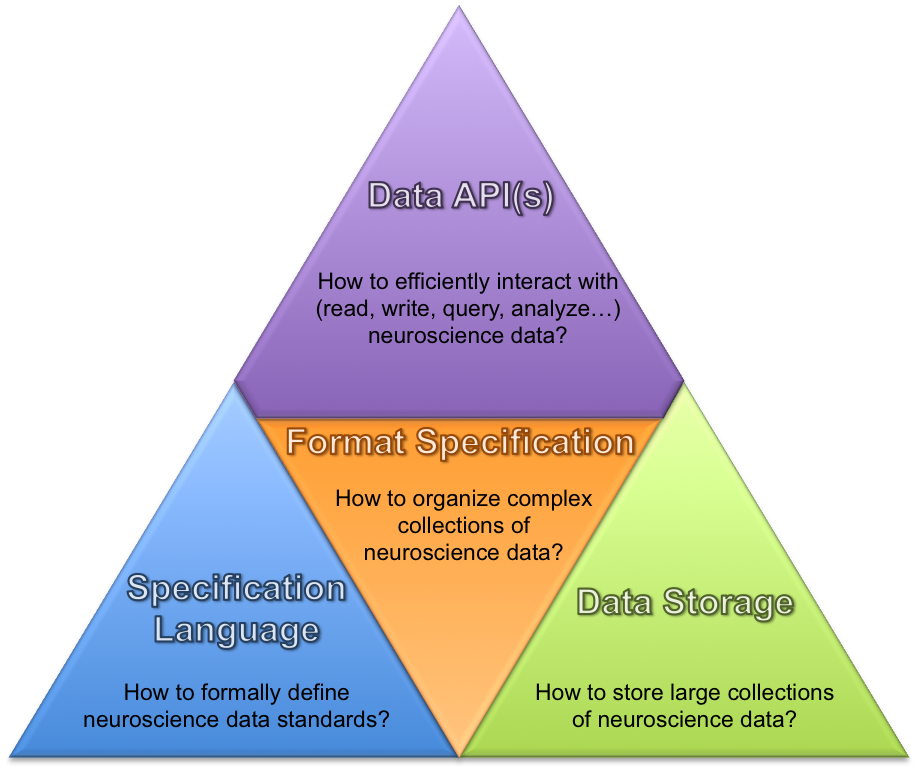
New modular software architecture and APIs to enable users/developers to efficiently interact with the…
News: Prominent U.S. Research Institutions Announce Collaboration Toward Sharing and Standardizing Neuroscience Data
The initiative is hoped to be a major step toward accelerating the pace of discoveries about the brain in health and disease.
The Allen Institute for Brain Science, California Institute of Technology, New York University School of Medicine, the Howard Hughes Medical Institute (HHMI) and the University of California, Berkeley (UC Berkeley) are collaborating on a project aimed at making databases about the brain more useable and accessible for neuroscientists – a step seen as critical to accelerating the pace of discoveries about the brain in health and disease. With funding from GE, The Kavli Foundation, the Allen Institute for Brain Science, the HHMI, and the International Neuroinformatics Coordinating Facility (INCF), the year-long project will focus on standardizing a subset of neuroscience data, making this research simpler for scientists to share.
This is the first collaboration launched by “Neurodata Without Borders,” a broader initiative with the goal of standardizing neuroscience data on an international scale, making it more easily sharable by researchers worldwide. This first project is called “Neurodata Without Borders: Neurophysiology.”
Unlike image file formats such as jpeg or tiff, that store digital information when we take a photo with our mobile phones and allow us to share that photo with anyone with a computer, no such data standard exists in neuroscience. However, developing such a standard, or unified data format, would enhance the ability of brain researchers worldwide to share and combine their research results. This would not only drive progress in neuroscience but also encourage the validation of existing results and create vital new collaborations with other fields.
“Neuroscientists aren’t limited by memory storage anymore; we’re limited by our ingenuity, the availability of data and our ability to talk to each other,” says Christof Koch, Chief Scientific Officer at the Allen Institute. “This pilot program is an effort to help us speak the same language.”
Today, researchers can simultaneously record the electrical or optical activity of a thousand neurons in a mouse’s brain while the animal is navigating a maze, for example, and that number may soon be in the millions. These recent, rapid technical advances mean that neuroscientists are generating data that is quantitatively and qualitatively different than before. But the languages, or formats, they use to capture those data (as well as the software tools they use to access and analyze them) vary from laboratory to laboratory-and sometimes even within a laboratory. This lack of uniformity makes it challenging to share and integrate experimental data-the raw material of science-and to mine and extract the most value from them.
The need for a common data format in neuroscience is made more urgent by the rise of large-scale collaborative projects, such as the Brain Research through Advancing Innovative Neurotechnologies (BRAIN) Initiative in the United States.
“These new initiatives are going to produce masses of data, but if it isn’t interchangeable and comparable, it’s just not going to be useful,” says Koch.
On a practical level, scientific publishers and granting agencies, such as the U.S. National Institutes of Health, are moving toward mandating data sharing as a requirement for funding.
“This is following on other efforts at openness in science,” says Markus Meister, a professor of biology at Caltech whose research group is supplying experimental data to the project. “The idea is that the material or resources that were developed with government funding or published in a journal have to be made available. But for neurophysiology data, there is no organized mechanism for doing that at the moment.”
The initial one-year program focuses on a subset of neuroscience data: cell-based neurophysiology data, which is sought-after by theorists who are building models of how the brain works. The partners will work with software developers and vendors to establish an open format that can store electrical and optical recordings of neural activity, and, importantly, the conditions under which an experiment was performed, such as how brain activity was recorded, how the animal was behaving at certain time points, and its species, sex and age. These “metadata” are often lost and yet without them the research results are meaningless.
This “metadata problem” poses an enormous challenge, says Friedrich Sommer, a theoretical neuroscientist at UC Berkeley who oversees an existing repository, CRCNS.org, where the neurophysiology datasets of Neurodata Without Borders will be stored and shared. UC Berkeley is coordinating Neurodata Without Borders with staff from the Allen Institute.
As Sommer explains, once a data format has been selected and extended, the neurophysiology datasets will be translated into the new common language and shared with the broader neuroscience community through the repository. Lastly, “application programming interfaces” (APIs) will be developed to allow researchers to use the common format for their own data with ease.
To get to that point, Neurodata Without Borders is calling on the neuroscience community to get involved. “We want to solicit the best ideas for the data format, so we are inviting researchers to look at the datasets which are now shared in their current format at CRCNS.org. Our hope is to engage the community to contribute ideas or propose their own data format for consideration,” says Sommer.
The most promising approaches to a common data format will be discussed, tested and extended at Neurodata Without Borders Hackathons, the first of which will be held in late November, to drive the rapid development of innovative software tools.
“The project has an aggressive timeline, but in a year’s time, the goal is to come up with a standard for neurophysiology data that we can agree on. We may not get it 100% right for 100% of researchers, but we’ll make a very good attempt,” says neuroscientist Karel Svoboda, group leader at HHMI’s Janelia Research Campus and a data-provider to Neurodata Without Borders. “Then, by buying into the data format ourselves-by explicitly moving our data into the format and making them available, we’ll set an example of how it could be done, and hopefully have others in the neuroscience community follow in our footsteps.”
There have been smaller efforts to develop a common language for neuroscience data in the past but they have fallen short of meeting the goals of the new project.
“This new effort is not very different in spirit from past ones, but it’s at a much larger scale,” says NYU School of Medicine’s Gyorgy Buzsaki, a pioneer of data-sharing in neuroscience and another data-provider. “We’re trying to make Neurodata Without Borders as attractive as possible by improving the quality of the datasets and how they are documented, and by thinking hard about how best to help researchers navigate around in them. Ultimately, if researchers can find and access the data they’re interested in in a few hours, they will choose it. But that is a very difficult thing to do.”
“With the emergence of large-scale brain initiatives around the world, data reuse and sharing becomes more important than ever. This project will facilitate neuroscience collaboration at a global scale,” says Sean Hill, Scientific Director, INCF.
“Standardizing a subset of neuroscience data is vital to accelerate the pace of research and innovation in brain health. We are proud to be working with these best-in-class organizations on such an important and needed study,” says Robert Wells, executive director, healthymagination strategy, GE.
Miyoung Chun, Executive Vice President of Science Programs, The Kavli Foundation, agrees with these assessments. “In neuroscience, as in many scientific fields, there are massive amounts of ‘Big Data’ but no coherent way to retrieve and use this information. Our hope is Neurodata Without Borders: Neurophysiology is a major step toward changing this and speeding breakthroughs in brain science.”